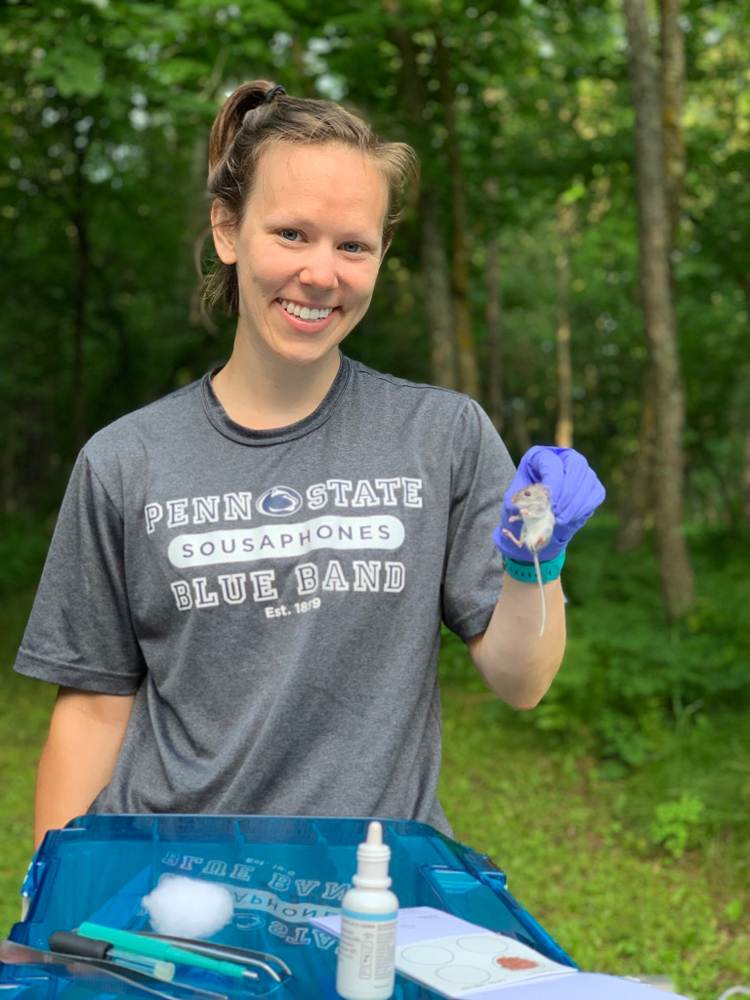
Janine Mistrick
Research Q & A
What’s your hometown?
State College, Pennsylvania
What are you currently working on?
Thanks to support provided by the Bell Museum graduate student awards through the Zoological Society Fund, this past summer (June–August 2019), I live-trapped wild Peromyscus mice around Minnesota and took blood and fecal samples to test for the presence or absence of various parasites: Giardia spp. and Cryptosporidium spp. in the feces (bacteria which cause diarrheal diseases in humans and livestock) and Borrelia burgdorferi in the blood (the causative agent of Lyme disease). I’m interested in whether there are geographic differences in disease prevalence (northern vs central Minnesota); differences in disease prevalence between mice found in more natural, wooded habitats or those found in/around buildings near people; and if there are genetic differences in the parasites on a geographic scale across the state or between natural vs. human-dominated habitats.
How are you working toward that goal?
All my samples from the summer are currently stored in freezers on the UMN Twin Cities campus in St. Paul. I’ve extracted DNA from the approximately 180 fecal samples I collected this summer and am in the process of screening those samples for Giardia intestinalis (the species of Giardia known to cause disease in humans). I will be sequencing positive samples as I find them to determine the genetic code of those samples and compare genetic diversity within and between geographic locations or sites with varying proximity to humans.
Why are you focusing your work in that area?
I am interested in wildlife diseases and how various aspects of disease (or the parasites that cause disease) vary across different habitats or geographic space. Working with genetic sequences of the parasites in this study also gives me valuable experience working with lab-based techniques, genetic sequencing protocols and technology, and computer programs to analyze and make sense of the genetic data. All of these are new, valuable skills in the field of disease ecology that will broaden my options for future research.
Where are you working on research/field work?
I trapped mice over the summer at Cedar Creek Ecosystem Science Reserve in East Bethel, Minnesota, and at Itasca State Park and the Itasca Biological Station and Laboratories in Clearwater County, Minnesota. Currently, I’m conducting my lab work in the lab of my mentor and collaborator, Dr. Peter Larsen in the College of Veterinary Medicine, on the UMN Twin Cities campus in St. Paul. I am also working with the UMN Genomics Center to Sanger sequence the positive samples I have isolated.
What will your next steps/research be?
My plan is to conclude screening my samples for Giardia and sequencing positive samples by the end of the spring 2020 semester. I hope to be able to compile and publish these results. Many other aspects of this project remain to be funded to completion, but there is interest within Dr. Larsen’s lab to use new genetic sequencing technology to rapidly screen the blood samples for presence/absence of Borrelia burgdorferi and/or to use the same technology to develop a rapid test to differentiate the two species of Peromyscus mice (P. leucopus and P. maniculatus) found in Minnesota (of which I may have representatives of both species) using genetic methods (these species are too similar to be visually differentiated in the field).
The research focused on my dissertation will be taking a new direction in summer 2020 when I will be traveling to Finland to work on a grant funded by the National Science Foundation among a team of researchers across universities in the U.S. which will be looking at wild vole populations and environmental factors which drive population-level patterns of hantavirus infection. While the study system will be changing, the field skills and lab techniques I have used with this project will continue to be useful and relevant to these future directions.